-Search query
-Search result
Showing 1 - 50 of 15,223 items for (author: ji & s)

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E
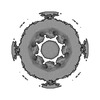
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-45206: 
Reconstituted P400 Subcomplex of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45240: 
P400 subcomplex of the native human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E

EMDB-45252: 
ARP module of the human TIP60 complex
Method: single particle / : Yang Z, Mameri A, Florez Ariza AJ, Cote J, Nogales E
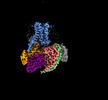
EMDB-60628: 
Carazolol-activated human beta3 adrenergic receptor
Method: single particle / : Zheng S, Zhang S, Dai S, Chen K, Gao K, Lin B, Liu X
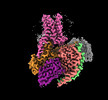
EMDB-60629: 
Epinephrine-activated human beta3 adrenergic receptor
Method: single particle / : Zheng S, Zhang S, Dai S, Chen K, Gao K, Lin B, Liu X

EMDB-45962: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18
Method: single particle / : Sun C, Jiang W

EMDB-45963: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18
Method: single particle / : Sun C, Jiang W

EMDB-45964: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated
Method: single particle / : Sun C, Jiang W

EMDB-45597: 
Cryo-EM structure of the human ether-a-go-go related K+ channel (hERG) in 300 mM K+
Method: single particle / : Lau CHY, Hunter MJ, Vandenberg JI

EMDB-45598: 
Cryo-EM structure of the human ether-a-go-go related K+ channel (hERG) in 3 mM K+
Method: single particle / : Lau CHY, Hunter MJ, Vandenberg JI

EMDB-45599: 
Cryo-EM structure of the human ether-a-go-go related K+ channel (hERG) in 300 mM K+ without symmetry
Method: single particle / : Lau CHY, Hunter MJ, Vandenberg JI

EMDB-45600: 
Cryo-EM structure of the human ether-a-go-go related K+ channel (hERG) in 3 mM K+ without symmetry
Method: single particle / : Lau CHY, Hunter MJ, Vandenberg JI

EMDB-38845: 
Icosahedrally averaged cryo-EM reconstruction of PhiKZ capsid before applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38846: 
Block 1 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-38848: 
Block 2 of PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-39002: 
Composite cryo-EM map of PhiKZ capsid after applying the "block-based" reconstruction method
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

PDB-8y6v: 
Near-atomic structure of icosahedrally averaged jumbo bacteriophage PhiKZ capsid
Method: single particle / : Yang Y, Shao Q, Guo M, Han L, Zhao X, Wang A, Li X, Wang B, Pan J, Chen Z, Fokine A, Sun L, Fang Q

EMDB-37571: 
cryo-EM structure of alligator haemoglobin in carbonmonoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37572: 
cryo-EM structure of alligator haemoglobin in oxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37573: 
cryo-EM structure of alligator haemoglobin in deoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37574: 
cryo-EM structure of human haemoglobin in carbonmonoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37575: 
cryo-EM structure of human haemoglobin in oxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

EMDB-37576: 
cryo-EM structure of human haemoglobin in deoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

PDB-8wix: 
cryo-EM structure of alligator haemoglobin in carbonmonoxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH

PDB-8wiy: 
cryo-EM structure of alligator haemoglobin in oxy form
Method: single particle / : Takahashi K, Lee Y, Fago A, Bautista NM, Kawamoto A, Kurisu G, Storz J, Nishizawa T, Tame JRH
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
